vedo

Your friendly python module
for scientific analysis and visualization of 3d objects.
💾 Installation
pip install vedo
additional installation details [click to expand]
- To install the latest _dev_ version of `vedo`: ```bash pip install -U git+https://github.com/marcomusy/vedo.git ``` - To install from the conda-forge channel: ```bash conda install -c conda-forge vedo ```🚀 Quick Start
from vedo import Sphere, show
sphere = Sphere().c("tomato")
show(sphere, axes=1).close()
This opens an interactive 3D window with a simple object and axes.
📙 Documentation
The webpage of the library with documentation is available here.
📌 Need help? Have a question, or wish to ask for a missing feature? Do not hesitate to ask any questions on the image.sc forum or by opening a github issue.
🎨 Features
The library includes hundreds of working examples for a wide range of functionalities
working with polygonal meshes and point clouds [click to expand]
- Import meshes from VTK format, STL, Wavefront OBJ, 3DS, Dolfin-XML, Neutral, GMSH, OFF, PCD (PointCloud), - Export meshes as ASCII or binary to VTK, STL, OBJ, PLY ... formats. - Analysis tools like Moving Least Squares, mesh morphing and more.. - Tools to visualize and edit meshes (cutting a mesh with another mesh, slicing, normalizing, moving vertex positions, etc..). - Split mesh based on surface connectivity. Extract the largest connected area. - Calculate areas, volumes, center of mass, average sizes etc. - Calculate vertex and face normals, curvatures, feature edges. Fill mesh holes. - Subdivide faces of a mesh, increasing the number of vertex points. Mesh simplification. - Coloring and thresholding of meshes based on associated scalar or vectorial data. - Point-surface operations: find nearest points, determine if a point lies inside or outside of a mesh. - Create primitive shapes: spheres, arrows, cubes, torus, ellipsoids... - Generate glyphs (associate a mesh to every vertex of a source mesh). - Create animations easily by just setting the position of the displayed objects in the 3D scene. Add trailing lines and shadows to moving objects is supported. - Straightforward support for multiple sync-ed or independent renderers in the same window. - Registration (alignment) of meshes with different techniques. - Mesh smoothing. - Delaunay triangulation in 2D and 3D. - Generate meshes by joining nearby lines in space. - Find the closest path from one point to another, traveling along the edges of a mesh. - Find the intersection of a mesh with lines, planes or other meshes. - Interpolate scalar and vectorial fields with Radial Basis Functions and Thin Plate Splines. - Add sliders and buttons to interact with the scene and the individual objects. - Visualization of tensors. - Analysis of Point Clouds - Moving Least Squares smoothing of 2D, 3D and 4D clouds - Fit lines, planes, spheres and ellipsoids in space - Identify outliers in a distribution of points - Decimate a cloud to a uniform distribution.working with volumetric data and tetrahedral meshes
- Import data from VTK format volumetric TIFF stacks, DICOM, SLC, MHD and more - Import 2D images as PNG, JPEG, BMP - Isosurfacing of volumes - Composite and maximum projection volumetric rendering - Generate volumetric signed-distance data from an input surface mesh - Probe volumes with lines and planes - Generate stream-lines and stream-tubes from vectorial fields - Slice and crop volumes - Support for other volumetric structures (structured and grid data)plotting and histogramming in 2D and 3D
- Polygonal 3D text rendering with Latex-like syntax and unicode characters, with 30 different fonts. - Fully customizable axis styles - donut plots and pie charts - Scatter plots in 2D and 3D - Surface function plotting - 1D customizable histograms - 2D hexagonal histograms - Polar plots, spherical plots and histogramming - Draw latex-formatted formulas in the rendering window. - Quiver, violin, whisker and stream-line plots - Graphical markers analogous to matplotlibintegration with other libraries
- Integration with the [Qt5](https://www.qt.io/) framework. - Interoperability with the [trimesh](https://trimsh.org/), [pyvista](https://github.com/pyvista/pyvista) and [pymeshlab](https://github.com/cnr-isti-vclab/PyMeshLab) libraries. - Export 3D scenes and embed them into a [web page](https://vedo.embl.es/examples/export_x3d.html). - Embed 3D scenes in *jupyter* notebooks with [K3D](https://github.com/K3D-tools/K3D-jupyter) (can export an interactive 3D-snapshot page [here](https://vedo.embl.es/examples/geo_scene.html)).⌨ Command Line Interface
Visualize a polygonal mesh or a volume from a terminal window simply with:
vedo https://vedo.embl.es/examples/data/embryo.tif
volumetric files (slc, tiff, DICOM...) can be visualized in different modes [click to expand]
|Volume 3D slicing`vedo --slicer embryo.slc`| Ray-casting
`vedo -g`| 2D slicing
`vedo --slicer2d`| |:--------|:-----|:--------| | 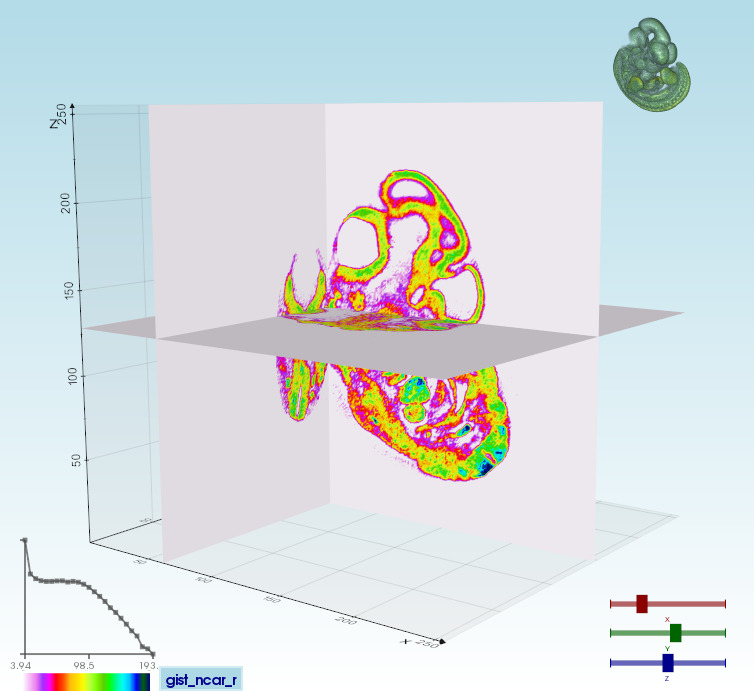 |  |  |
Type vedo -h for the complete list of options.
🐾 Gallery
vedo currently includes hundreds of working examples and notebooks.
Run any of the built-in examples. In a terminal type: vedo -r warp2
Check out the example galleries organized by subject here:

</a>
✏ Contributing
Any contributions are greatly appreciated. If you have a suggestion, bugfix, feature, or documentation improvement, please open an issue or submit a pull request.
See CONTRIBUTING.md for contribution guidelines and workflow details.
📜 References
Scientific publications leveraging vedo:
browse 68 publications using vedo [click to expand]
**2026**
- L. Aviñó-Esteban *et al.*, *"Limblab: pipeline for 3D analysis and visualisation of limb bud gene expression"*, BMC Bioinformatics 27(1): 6 (2026).
- D. Krsikapa, I. Y. Kim, *"Gradient-based optimization of component layout: addressing accessibility and mounting in assembly system design"*, Journal of Mechanical Design 148(3): 031702 (2026).
**2025**
- A. Kharlamova *et al.*, *"Spatial CAPTCHA: Generatively Benchmarking Spatial Reasoning for Human-Machine Differentiation"*, arXiv preprint arXiv:2510.03863 (2025).
- J. F. Fuhrmann *et al.*, *"Apical extracellular matrix regulates fold morphogenesis in the Drosophila wing disc"*, bioRxiv 2025-09 (2025).
- B. Li *et al.*, *"Three-dimensional spatial transcriptomics at isotropic resolution enabled by generative deep learning"*, bioRxiv 2025-08 (2025).
- T.-T. Hsu *et al.*, *"Shared Alteration of Whole-Brain Connectivity and Olfactory Deficits in Multiple Autism Mouse Models"*, bioRxiv 2025-02 (2025).
- A. Arrabi *et al.*, *"C-arm guidance: A self-supervised approach to automated positioning during stroke thrombectomy"*, 2025 IEEE 22nd International Symposium on Biomedical Imaging (ISBI).
- L. Aviñó-Esteban, H. Cardona-Blaya, J. Sharpe, *"Spatio-temporal reconstruction of gene expression patterns in developing mice"*, Development 152: DEV204313 (2025), [DOI](https://doi.org/10.1242/dev.204313).
- B. Bortolon *et al.*, *"GRASPLAT: Enabling dexterous grasping through novel view synthesis"*, 2025 IEEE/RSJ International Conference on Intelligent Robots and Systems (IROS).
- L. Carreira *et al.*, *"Targeted nano-energetic material exploration through active learning algorithm implementation"*, Energetic Materials Frontiers 6(1): 3-13 (2025).
- M. Chirillo *et al.*, *"PyReconstruct: A fully open-source, collaborative successor to Reconstruct"*, Proceedings of the National Academy of Sciences 122(31): e2505822122 (2025).
- B. Clayton *et al.*, *"A facile method to create continuum stochastic sheet-based cellular materials"*, Additive Manufacturing: 104917 (2025).
- A. Gross *et al.*, *"STRESS, an automated geometrical characterization of deformable particles for in vivo measurements of cell and tissue mechanical stresses"*, Scientific Reports 15(1): 28599 (2025).
- A. Gauvain *et al.*, *"HydroModPy: A Python toolbox for deploying catchment-scale shallow groundwater models"* (2025).
- K. N. Halwachs *et al.*, *"Effects of Stiffness and Degradability on Cardiac Fibroblast Contractility and Extracellular Matrix Secretion in Three-Dimensional Hydrogel Scaffolds"*, ACS Biomaterials Science & Engineering 11(11): 6521-6533 (2025).
- R. Kliman *et al.*, *"Toward an Automated System for Nondestructive Estimation of Plant Biomass"*, Plant Direct 9(3): e70043 (2025).
- J. Laussu *et al.*, *"Deciphering the interplay between biology and physics with a finite element method-implemented vertex organoid model: A tool for the mechanical analysis of cell behavior on a spherical organoid shell"*, PLOS Computational Biology 21(1): e1012681 (2025).
- M. Mitelut *et al.*, *"Continuous monitoring and machine vision reveals that developing gerbils exhibit structured social behaviors prior to the emergence of autonomy"*, PLoS Biology 23(9): e3003348 (2025).
- J.S. Posada *et al.*, *"morphoHeart: A quantitative tool for integrated 3D morphometric analyses of heart and ECM during embryonic development"*, PLOS Biology 23(1) (2025), [DOI](https://doi.org/10.1371/journal.pbio.3002995).
- A. Prashanth, S. Hathwar, *"Comparing the Effectiveness of Deep Learning Models Combined with Loss Functions in Cardiac Segmentation"* (2025).
- M. Levin Thomas *et al.*, *"Banner cloud formation at the Matterhorn: Measurements versus large-eddy simulations"*, Journal of the Atmospheric Sciences 82(8): 1661-1675 (2025).
- H. Xu, *"A Progressive Interactive Exploration Framework for Vector Field Data Guided by Storylines"*, 2025 18th International Congress on Image and Signal Processing, BioMedical Engineering and Informatics (CISP-BMEI).
- S. M. Zahedi *et al.*, *"Comparative evaluation of neural networks and transfer learning for predicting mechanical properties of 3D-printed bone scaffolds"*, Macromolecular Materials and Engineering 310(10): e00073 (2025).
**2024**
- C. Lei *et al.*, *"Automatic tooth arrangement with joint features of point and mesh representations via diffusion probabilistic models"*, Computer Aided Geometric Design 111: 102293 (2024), [Code](https://github.com/lcshhh/TADPM).
- S. Li *et al.*, *"MogaNet: Multi-order Gated Aggregation Network"*, International Conference on Learning Representations (2024).
- J. Cotterell *et al.*, *"Cell 3D Positioning by Optical encoding (C3PO) and its application to spatial transcriptomics"*, bioRxiv 2024.03.12.584578 (2024), [DOI](https://doi.org/10.1101/2024.03.12.584578).
- D. Galvez Alcantara, *"Development of a finite element framework for biological applications"* (2024).
- M. Gazziro *et al.*, *"Fully Automated Ultra-Personalized 3D Printed Prosthetic Breasts"*, American Journal of Biomedical Science & Research 20: 128-132 (2024).
- I. G. Gonçalves, J. M. García-Aznar, *"Neurorosettes: a novel computational modelling framework to investigate the Homer-Wright rosette formation in neuroblastoma"*, Computational Particle Mechanics 11(2): 565-577 (2024).
- E. Guiltinan *et al.*, *"pySimFrac: A Python library for synthetic fracture generation and analysis"*, Computers & Geosciences 191: 105665 (2024).
- R. Haase *et al.*, *"Benchmarking large language models for bio-image analysis code generation"*, bioRxiv 2024-04 (2024).
- Y. Jiang, S. L. Bugby, J. E. Lees, *"PMST: A custom Python-based Monte Carlo Simulation Tool for research and system development in portable pinhole gamma cameras"*, Nuclear Instruments and Methods in Physics Research Section A 1061: 169161 (2024).
- D. Li, F. Pucci, M. Rooman, *"Prediction of paratope-epitope pairs using convolutional neural networks"*, International Journal of Molecular Sciences 25(10): 5434 (2024).
- M. Marro, L. Moccozet, D. Vernez, *"A numerical model for quantifying exposure to natural and artificial light in human health research"*, Computers in Biology and Medicine 171: 108119 (2024).
- M. Deepa Maheshvare *et al.*, *"Kiphynet: an online network simulation tool connecting cellular kinetics and physiological transport"*, Metabolomics 20(5): 94 (2024).
- S. Scholz *et al.*, *"Factors influencing pain medication and opioid use in patients with musculoskeletal injuries: a retrospective insurance claims database study"*, Scientific Reports 14(1): 1978 (2024).
- J. Sultana, M. Naznin, T. R. Faisal, *"SSDL - an automated semi-supervised deep learning approach for patient-specific 3D reconstruction of proximal femur from QCT images"*, Medical & Biological Engineering & Computing 62(5): 1409-1425 (2024).
- S. Wang *et al.*, *"A 3D dental model dataset with pre/post-orthodontic treatment for automatic tooth alignment"*, Scientific Data 11(1): 1277 (2024).
**2023**
- S. Baumer *et al.*, *"Robocasting of ceramic Fischer-Koch S scaffolds for bone tissue engineering"*, Journal of Functional Biomaterials 14(5): 251 (2023).
- R. Blain *et al.*, *"A tridimensional atlas of the developing human head"*, Cell 186(26): 5910-5924 (2023).
- B. Bogusławski *et al.*, *"Increasing brightness in multiphoton microscopy with a low-repetition-rate, wavelength-tunable femtosecond fiber laser"*, Optics Continuum 3(1): 22-35 (2023).
- G. Gust *et al.*, *"3D Analytics: Opportunities and Guidelines for Information Systems Research"*, arXiv preprint arXiv:2308.08560 (2023).
- T.-T. Hsu, C.-Y. Wang, Y.-P. Hsueh, *"Tbr1 autism mouse model displays altered structural and functional amygdalar connectivity and abnormal whole-brain synchronization"*, bioRxiv 2023-07 (2023).
- J. Laussu *et al.*, *"Deciphering interplay between biology and physics: finite element method-implemented vertex organoid model raises the challenge"*, bioRxiv 2023-05 (2023).
- Y. Li *et al.*, *"Research on the evolutionary history of the morphological structure of cotton seeds: a new perspective based on high-resolution micro-CT technology"*, Frontiers in Plant Science 14: 1219476 (2023).
- S. Monji-Azad *et al.*, *"SimTool: A toolset for soft body simulation using Flex and Unreal Engine"*, Software Impacts 17: 100521 (2023).
- S. Triarjo *et al.*, *"Automatic 3D digital dental landmark based on point transformation weight"*, 2023 International Conference on Artificial Intelligence in Information and Communication (ICAIIC).
- V. Zinchenko *et al.*, *"MorphoFeatures for unsupervised exploration of cell types, tissues, and organs in volume electron microscopy"*, eLife 12: e80918 (2023).
**2022**
- M. Blanc *et al.*, *"A dynamic and expandable digital 3D-atlas maker for monitoring the temporal changes in tissue growth during hindbrain morphogenesis"*, eLife 11: e78300 (2022).
- G. Dalmasso *et al.*, *"4D reconstruction of murine developmental trajectories using spherical harmonics"*, Developmental Cell 57, 1-11 September 2022, [DOI](https://doi.org/10.1016/j.devcel.2022.08.005).
- M. Deepa Maheshvare *et al.*, *"A Graph-Based Framework for Multiscale Modeling of Physiological Transport"*, Frontiers in Network Physiology 1: 802881 (2022), [DOI](https://www.frontiersin.org/articles/10.3389/fnetp.2021.802881/full).
- M. Erber *et al.*, *"Geometry-based assurance of directional solidification for complex topology-optimized castings using the medial axis transform"*, Computer-Aided Design 152: 103394 (2022).
- J. Hellar *et al.*, *"Manifold approximating graph interpolation of cardiac local activation time"*, IEEE Transactions on Biomedical Engineering 69(10): 3253-3264 (2022).
- A. Jaeschke, H. Eckert, L. J. Bray, *"Qiber3D - an open-source software package for the quantitative analysis of networks from 3D image stacks"*, GigaScience 11: giab091 (2022).
- J. Klatzow, G. Dalmasso, N. Martínez-Abadías, J. Sharpe, V. Uhlmann, *"µMatch: 3D shape correspondence for microscopy data"*, Frontiers in Computer Science (2022), [DOI](https://doi.org/10.3389/fcomp.2022.777615).
- N. Lamb *et al.*, *"DeepJoin: Learning a Joint Occupancy, Signed Distance, and Normal Field Function for Shape Repair"*, ACM Transactions on Graphics 41(6) (2022), [DOI](https://dl.acm.org/doi/abs/10.1145/3550454.3555470).
- J. E. Santos *et al.*, *"MPLBM-UT: Multiphase LBM library for permeable media analysis"*, SoftwareX 18: 101097 (2022).
- D. J. E. Waibel *et al.*, *"Capturing Shape Information with Multi-scale Topological Loss Terms for 3D Reconstruction"*, Lecture Notes in Computer Science 13434 (2022), [DOI](https://doi.org/10.1007/978-3-031-16440-8_15).
**2021**
- F. Claudi, A. L. Tyson, T. Branco, *"Brainrender. A python based software for visualisation of neuroanatomical and morphological data."*, eLife 10: e65751 (2021), [DOI](https://doi.org/10.7554/eLife.65751).
- F. Claudi, T. Branco, *"Differential geometry methods for constructing manifold-targeted recurrent neural networks"*, bioRxiv 2021.10.07.463479 (2021), [DOI](https://doi.org/10.1101/2021.10.07.463479).
- X. Lu *et al.*, *"3D electromagnetic modeling of graphitic faults in the Athabasca Basin using a finite-volume time-domain approach with unstructured grids"*, Geophysics (2021), [DOI](https://doi.org/10.1190/geo2020-0657.1).
- S. Ortiz-Laverde *et al.*, *"Proposal of an open-source computational toolbox for solving PDEs in the context of chemical reaction engineering using FEniCS and complementary components"*, Heliyon 7(1) (2021).
- J. Paglia *et al.*, *"TRACER: a toolkit to register and visualize anatomical coordinates in the rat brain"*, bioRxiv 2021-10 (2021).
- A. Pollack *et al.*, *"Stochastic inversion of gravity, magnetic, tracer, lithology, and fault data for geologically realistic structural models: Patua Geothermal Field case study"*, Geothermics 95 (2021), [DOI](https://doi.org/10.1016/j.geothermics.2021.102129).
**2020**
- J. S. Bennett, D. Sijacki, *"Resolving shocks and filaments in galaxy formation simulations: effects on gas properties and star formation in the circumgalactic medium"*, Monthly Notices of the Royal Astronomical Society 499(1) (2020), [DOI](https://doi.org/10.1093/mnras/staa2835).
- J. D. P. Deshapriya *et al.*, *"Spectral analysis of craters on (101955) Bennu"*, Icarus (2020), [DOI](https://doi.org/10.1016/j.icarus.2020.114252).
**2018**
- X. Diego *et al.*, *"Key features of Turing systems are determined purely by network topology"*, Physical Review X 8, 021071 (2018), [DOI](https://journals.aps.org/prx/abstract/10.1103/PhysRevX.8.021071).
- M. Musy, K. Flaherty *et al.*, *"A Quantitative Method for Staging Mouse Limb Embryos based on Limb Morphometry"*, Development 145(7): dev154856 (2018), [DOI](http://dev.biologists.org/content/145/7/dev154856).
Have you found this software useful for your research? Star ✨ the project and cite it as:
M. Musy et al.,
“vedo, a python module for scientific analysis and visualization of 3D objects and point clouds”,
Zenodo, 2021, doi: 10.5281/zenodo.7019968.
| |
| |
|:——–|:—–|

